How to download InterPro data?¶
InterPro data and search tools are freely available for download. We provide bulk downloads, data exports on each relevant InterPro page and an API to allow easy access for user scripts.
Download page¶
This is available under the Download section in the navigation menu. This page is divided into multiple tabs.
The InterPro tab provides various files containing pre-calculated InterPro data for the current release that can be downloaded. Data from previous releases are available in the InterPro ftp.
The InterProScan tab provides a downloadable version of the InterProScan software.
The Pfam tab gives access various Pfam files. Data from previous Pfam releases are available in the Pfam ftp.
The PRINTS and SFLD tabs give access to the latest PRINTS and SFLD release files, available from the InterPro ftp.
The AntiFam tab give access to the latest AntiFam release files.
Export button¶
The export button, found on various entry pages in InterPro, is located next to the text filter at the top of result tables. It allows data to be downloaded as JSON or Tab Separated Values (TSV). The data sent from the InterPro Application Programming Interface (API) to populate the table can also be viewed using this component. When the file to generate is too big (bigger than 10K entities) we recommend to use the Script generator described in the section below to get the information from the API.
Your downloads¶
This page is accessible through the Results tab in the navigation menu, under “Your downloads” section.
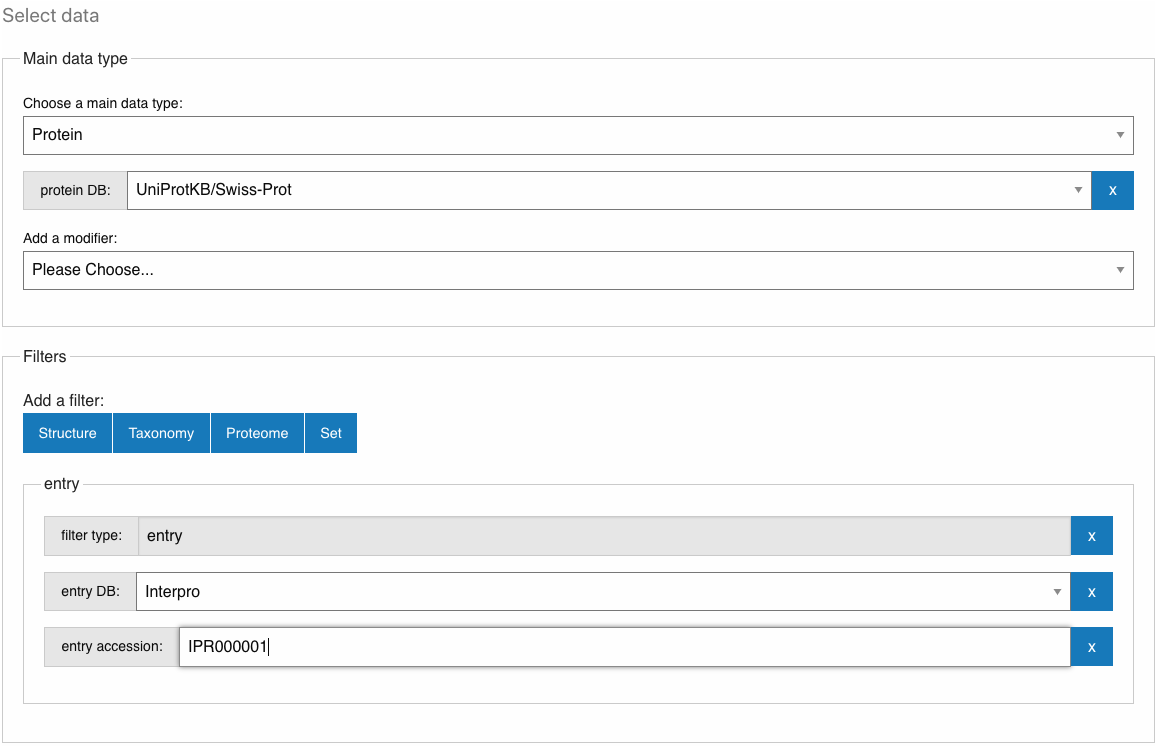
The purpose of this page is to give the user a way to select and filter InterPro data. Filtered data can then be downloaded in different file formats (if the selection has less than 10K entities), using the provided API call or through a script generated automatically.
For Example, the image above shows Protein as the main data type selected and it will only select proteins included in the database UniProtKB/Swiss-Prot; this selection is then filtered by the selection of the endpoint entry with InterPro as the database and accession IPR000001. In other words this will generate the list of SwissProt proteins that are matching IPR000001 (also available under the Proteins tab in the InterPro entry page for IPR000001, with the reviewed option selected). The results are stored in the browser (IndexedDB), allowing to retrieve previous searches.
Output formats¶
The following output formats are currently supported, if the number of entities selected is lower than 10K:
Text: a list of accessions, 1 per line
FASTA: a single file with multiple sequences in Fasta format (only available for proteins)
JSON: it reuses the format returned by the InterPro API.
TSV: reformats the JSON from the API to create a TSV file.
After selecting the output format, clicking on the Download button at the bottom of the page will start the downloading.
Programming scripts¶
The script can be generated in 4 different languages: Python 3, Perl and JavaScript, it allows the download of the filtered data directly from the InterPro API and can be integrated in the users own program.
InterPro Application Programming Interface (API)¶
The InterPro API provides programmatic access to all the InterPro entries and their related entities in Json format.The API has six main endpoints, which corresponds to the InterPro data types: entry, protein, structure, taxonomy, proteome and set.
An API call is formed of one or multiple endpoint blocks. An endpoint block consists of a data type, a source database and an accession (e.g. api/datatype/sourcedb/accession).
For example the URL /entry/interpro provides a pageable list of all the InterPro entries. And the URL /protein/uniprot/p99999 returns all the details of the protein identified with the UniProt accession P99999.
The combined URL /entry/interpro/protein/uniprot/p99999 returns the list of all the InterPro entries that match in the P99999 protein accession.
For more information on how to use the InterPro API, you can watch this recorded webinar or continue with the guide below.